There are several companies that offer a consumer genome test to analyze your ancestry, and some cases, health traits as well. These include 23andMe, Ancestry.com, and FTDNA.com.
Each of these companies allows you to download the raw data from the genome test. You can use this raw data to analyze health information with third-party software, like Enlis Genome Personal.
Which company has the most health information in their raw data? Let’s find out!
Note: 23andMe recently revamped their online service, but in the documentation, they state that the genotyping chip has not changed. The v4 chip, launched in December 2013, is still being used.
We used the Enlis import website to import recent raw genome data from each of 23andMe, Ancestry.com, and FTDNA.com. A PDF summary report is generated for each genome, and delivered by email. This summary report includes a section that describes how many disease or trait positions were successfully sequenced. (Known Phenotype Summary) (See an example report here)
Our current database contains 42,032 genome positions linked to a disease or trait.
The results from each company are as follows:
| 23andMe.com (v4) |
Ancestry.com | FTDNA.com | |
| Disease or trait positions successfully sequenced | 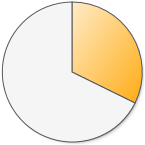 13,537 (32.2%) 13,537 (32.2%) |
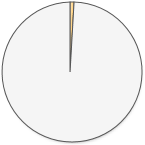 417 (1.0%) 417 (1.0%) |
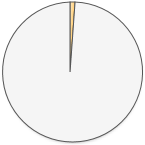 472 (1.1%) 472 (1.1%) |
| Missing data positions | 28,495 (67.8%) | 41,615 (99.0%) | 41,560 (98.9%) |
There is one clear winner here! 23andMe’s raw data has by far the most disease and trait information. When 23andMe designed their genotyping chip, they focused on adding SNPs that are already known to be involved in health. Here is a complete list of the diseases and traits found in 23andMe’s raw data.
Comparing 23andMe chip versions
23andMe has revised their genotyping chips several times over the past few years. Here we compare version 2, version 3, and the most recent, version 4.
| 23andMe.com (v2) |
23andMe.com (v3) | 23andMe.com (v4) | |
| Number of SNPs on chip | ~576,000 | ~967,000 | ~602,000 |
| Disease or trait positions successfully sequenced | 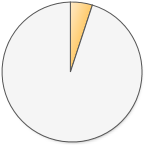 2,157 (5.1%) 2,157 (5.1%) |
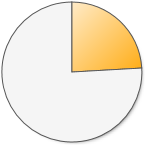 10,101 (24.0%) 10,101 (24.0%) |
 13,537 (32.2%) 13,537 (32.2%) |
| Missing data positions | 39,875 (94.9%) | 31,931 (76.0%) | 28,495 (67.8%) |
With every version, 23andMe is adding more disease and trait SNPs. It’s interesting to note that although the v4 chip tested fewer SNPs overall (compared to v3), it did increase the number of disease and trait SNPs tested.
Comparing 23andMe to next-generation sequencing
23andMe’s genotyping service doesn’t sequence your entire genome, only very select parts of the genome. With the newer next-generation sequencing technology, we can get exome or whole genome data. Exome data includes only about 3% of the entire genome, but it’s the part that we know the most about, and the part that many suspect is the most important. So how does 23andMe compare to exome or whole genome data?
| 23andMe.com (v4) |
Exome | Whole genome | |
| Percent of genome sequenced | 0.02% | 3-5% | 90-95% |
| Disease or trait positions successfully sequenced |  13,537 (32.2%) 13,537 (32.2%) |
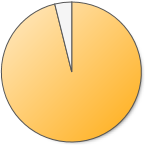 40,405 (96.1%) 40,405 (96.1%) |
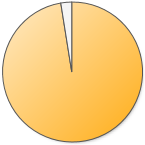 40,979 (97.5%) 40,979 (97.5%) |
| Missing data positions | 28,495 (67.8%) | 1,627 (3.9%) | 1,053 (2.5%) |
Next-generation sequencing data has a large advantage over 23andMe data and this gap will only widen as we learn more about the human genome.
Recommendations:
- Â If you are only interested in ancestry information, any of the 3 consumer services above will do.
- Â If you want ancestry information with some additional health information, 23andMe is the best.
- Â If you are most interested in known health information, or are looking for a unknown cause of a disease or trait, an exome is the most cost effective solution, while a whole genome sequence is the most comprehensive and future proof.



